Cell types: from genes to circuits
Part II in our series on cell types and connectomes
In the last post, I presented a counterintuitive idea: areas of the brain that contain many cell types encode innate behaviors and rewards. Myriad neurons with strange morphology that are connected idiosyncratically are an indicator of evolutionary pressure: behaviors and rewards that have survival value become genetically hard-coded. What I didn’t get to is how, mechanically, cell types can encode bespoke circuits. How is it that two neurons of different types can “decide” whether or not to connect to each other? And how can that encode important circuit motifs, like point and line attractors?
It turns out that evolution has (at least) two different tricks to encode intrinsic circuits: wiring rules and wireless (neuropeptide) connection rules. I present how these different rules interact in a circuit in the hypothalamus that controls aggression. Let’s go!
Cell Types Determine Wiring Instructions
Dániel and Albert-László Barabási propose a formal framework for understanding how genetic identity translates into connectivity (Barabási & Barabási, 2019). In their model, each neuron possesses a “barcode”—a combinatorial pattern of transcription factors that defines its cell type. Wiring rules then operate on these barcodes through what they call biclique operators.
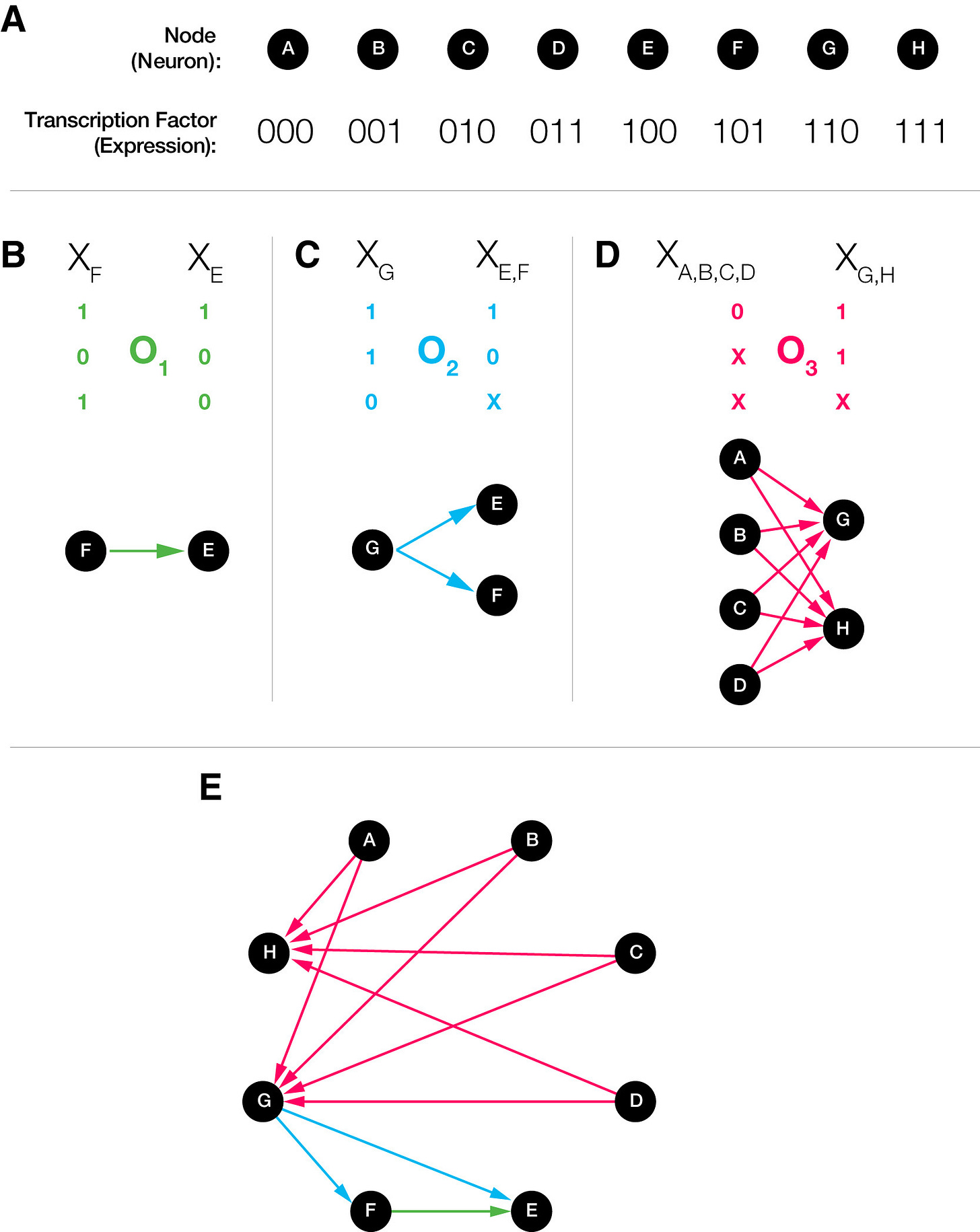
A biclique operator recognizes matching patterns in source and destination neurons and generates connections between all neurons matching those patterns. Consider a system where neurons are identified by three binary transcription factors. An operator might say “connect all neurons expressing TF1 and TF3 to all neurons expressing TF2 and TF3”. The key insight is that a single compact genetic rule can specify many connections simultaneously. Anything from OR gates to AND & NOR gates, combined with Boolean masks, can be expressed this way.
The Barabásis tested this model against the C. elegans connectome. The key test is that the pattern of connections in this model is highly non-random: cells of the same type should connect preferentially to the same target cell types. They found these patterns—biclique motifs—at a far higher rate than what random or models would predict. Moreover, neurons within these bicliques shared statistically significant gene expression patterns, suggesting that common genetic factors indeed drive synapse formation within these subgraphs. Similar patterns appeared in C. intestinalis and drosophila olfactory circuits.
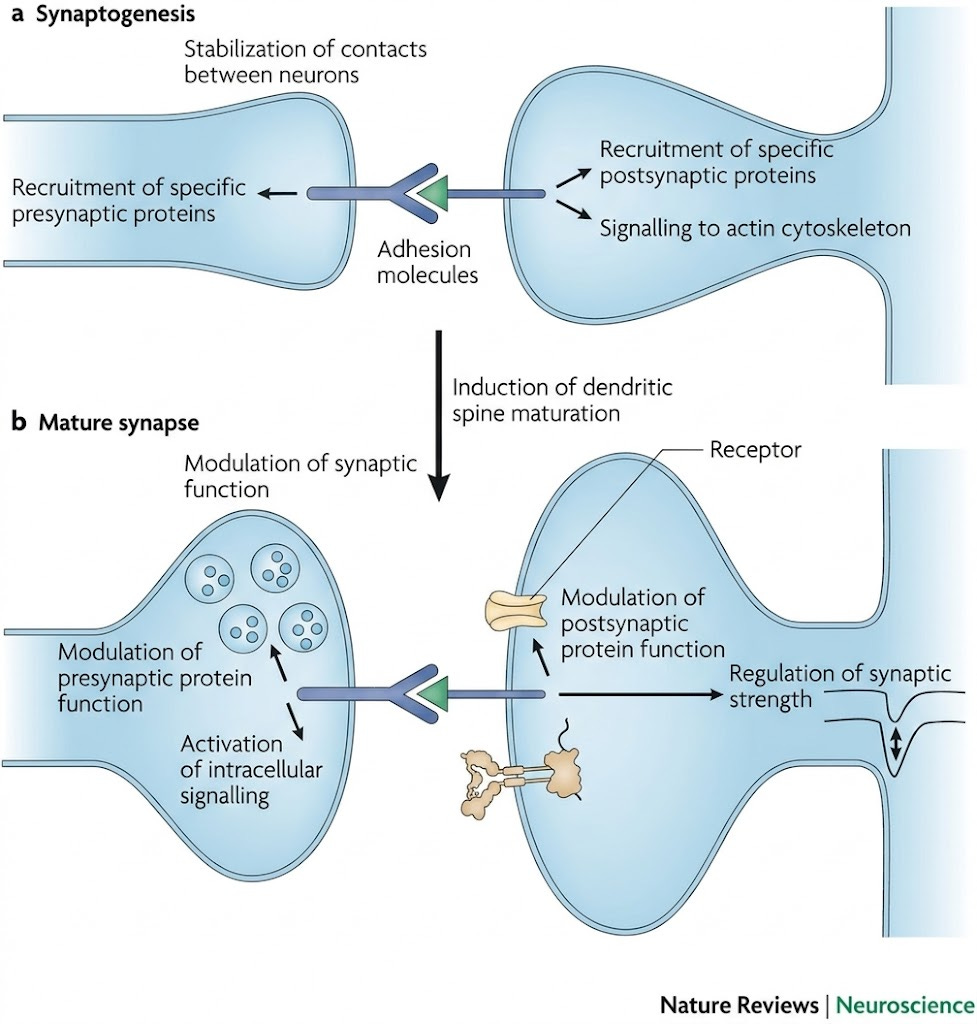
The concrete mechanism for biclique matching is not resolved in this computational paper. However, the discussion hints that it could involve cell adhesion molecules (CAMs) and receptors. When neurons of different types come into contact during development, surface proteins recognize each other through receptor-ligand interactions. If neurons of type A express receptor X and neurons of type B express ligand Y, and the X-Y combination triggers synapse formation, then all A will end up connecting with B (provided they’re within range).
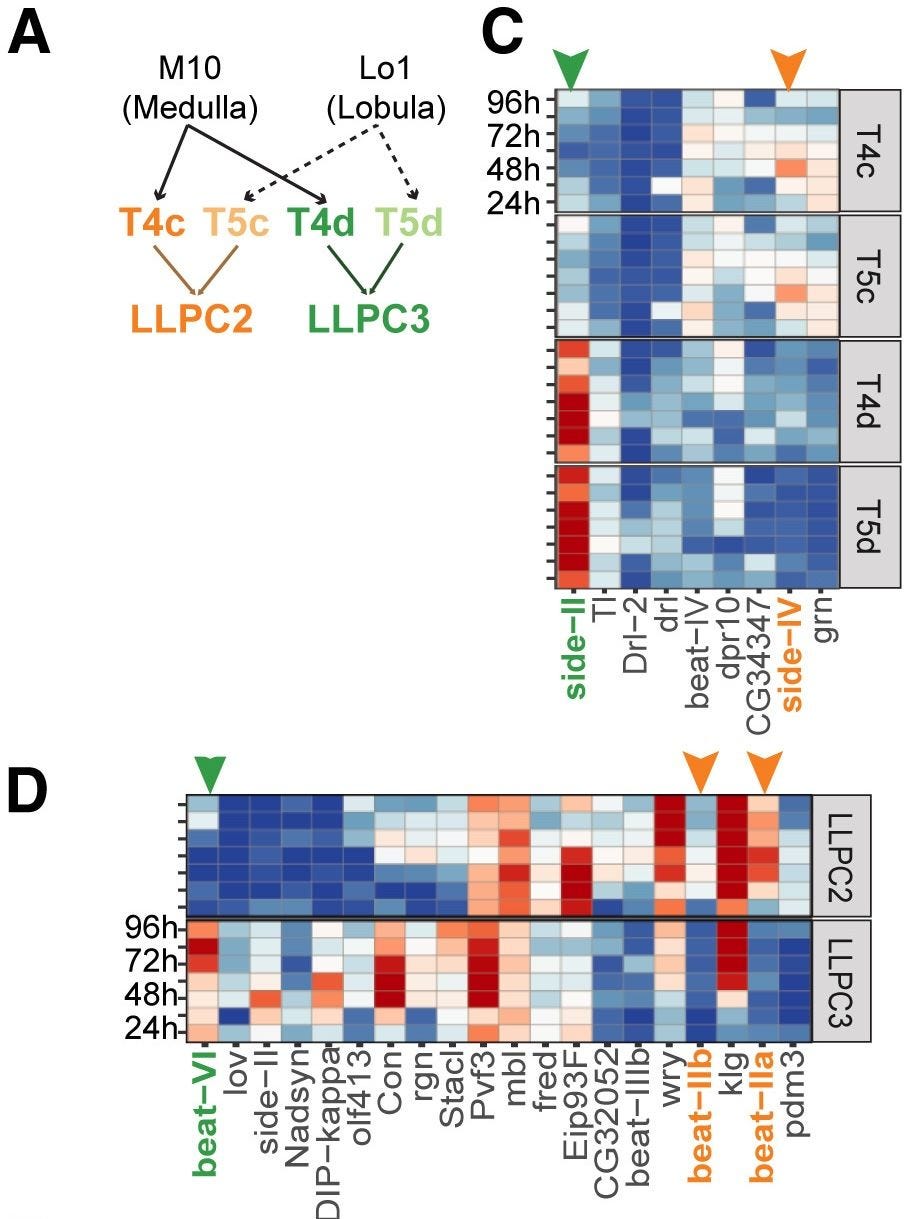
Indeed, a long line of work finds that CAMs allow different neuron types to very precisely find each other, even through the clutter of the neuropil. There’s a great paper from Yoo et al. (2023) that finds conceptually similar wiring rules in the fly’s visual system, where one has access to both transcriptomes and connectomes. Here, it’s the Side and Beat receptor-ligand pairs that determine with great precision how different motion-selective neurons connect. This is one way evolution writes circuit diagrams into the genome: by specifying the molecular handshakes that create connectivity patterns.
Connectivity Is Function
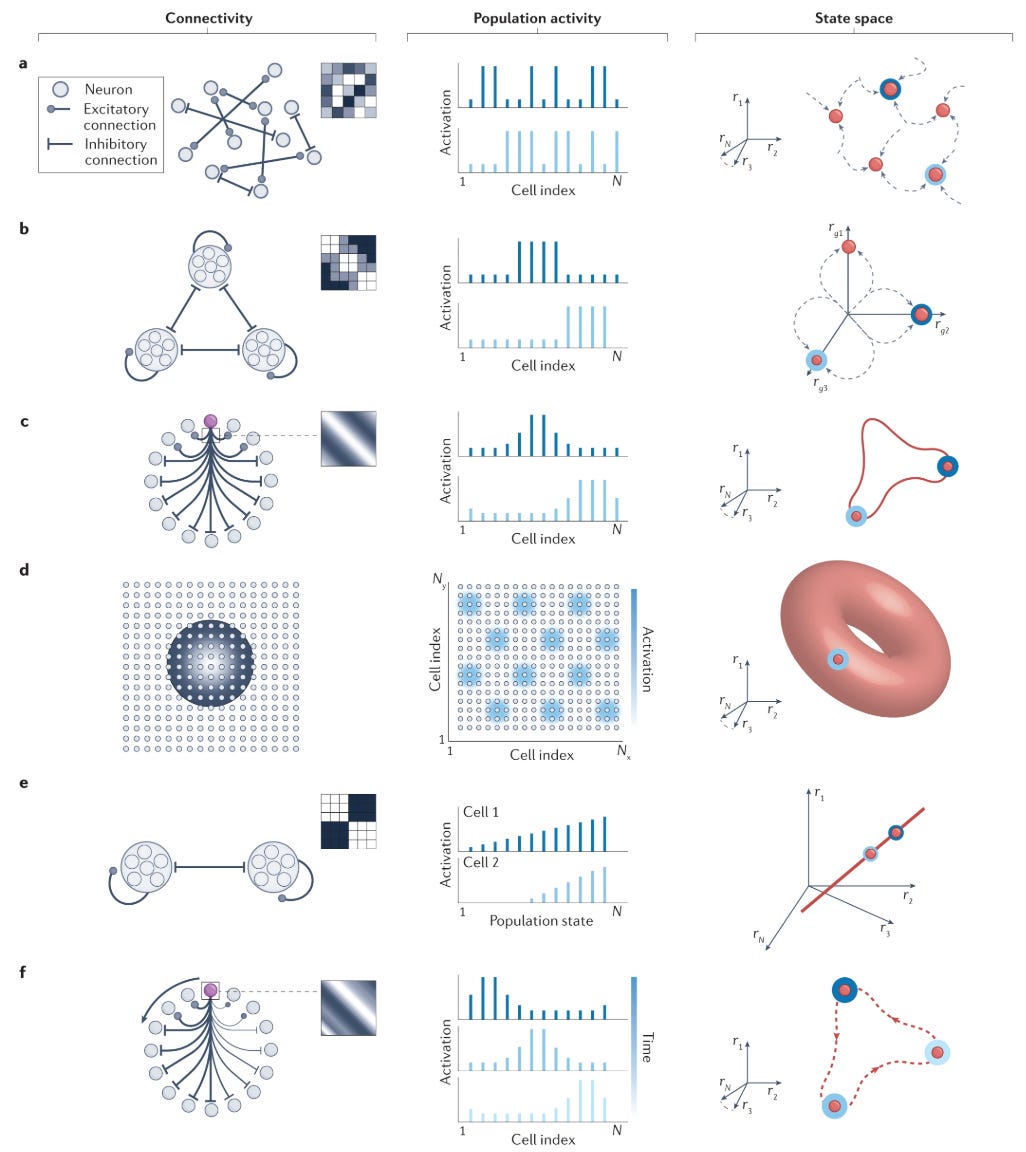
A deeper principle underlies the cell-type framework: connection topology determines computational function. By wiring up neurons in just the right way, we can create integrator, differentiators, and all sorts of bespoke circuits. A review by Khona and Fiete catalogs how different connectivity motifs produce different attractor dynamics (Khona & Fiete, 2022):
Point attractors arise from recurrent excitation balanced by global inhibition, implementing memory systems like the Hopfield network.
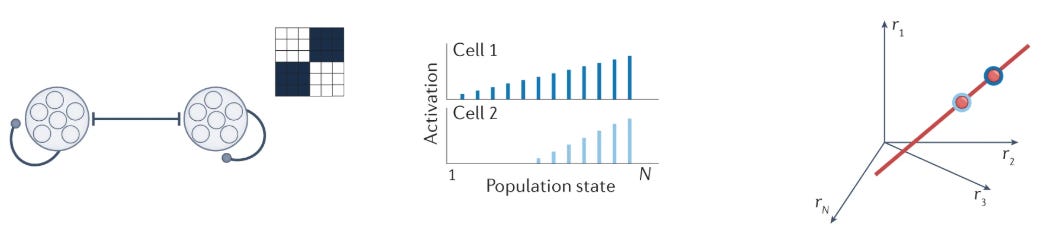
Line attractors—continuous manifolds of stable states—require precisely tuned excitation-inhibition balance with quasi-linear neural responses, enabling integrator circuits.
Ring attractors emerge when neurons form cyclic connectivity patterns, allowing, for example, head direction cells to encode angular position continuously.
Winner-take-all dynamics emerge from mutual excitation within groups combined with cross-group inhibition.
The implication is profound: by specifying which cell types connect to which—via CAM operators in the Barabási model—the genome encodes not just anatomy but dynamics. A circuit that integrates, oscillates, or makes winner-take-all decisions does so because its connection topology was specified to produce those dynamics.
The Wireless Connectome: Neuropeptides as Parallel Channels
The Barabási model describes the wired connectome—synaptic connections determined by cell adhesion molecules. But there is a parallel system equally important for genome-specified communication: neuropeptides, which constitute what we might call the wireless connectome. Some neuropeptides are household names, like oxytocin (often referred to in the popular media as the love hormone, although it’s much more complicated), glucagon-like peptide-1 (GLP-1, the receptor for blockbuster weight-loss drugs like Ozempic), or β-Endorphin. Others may be familiar to neurophysiologists, like SST and VIP, used as markers for subtypes of inhibitory cortical neurons.
When I first started learning about neuropeptides, it broke my mental model of how neurons work. Like neurotransmitters, neuropeptides come in pairs: ligands (encoded as neuropeptide precursors in the genome, NPPs) and receptors (G-protein coupled receptors, or GPCRs). However:
They don’t get released at synapses. They get packaged up in dense core vesicles and are released from the cell body or from axons.
They diffuse willy-nilly in the extracellular space through Brownian motion. Hence, “wireless transmission”. Their effective range is a few hundred micron radius ball around the diffusion location, though that can vary by neuropeptide species.
They are not tightly recycled or degraded like neurotransmitters. They can persist over minutes.
Sensitivity at the receptors is sky-high: in the nano-molar range.
There’s O(100) neuropeptide systems in the brain, covering, between protopeptides and receptors, upwards of 500 genes—about 2% of the human genome.
They’ve been around forever, predating neurons.
Whenever you think that the brain is really complicated, it usually turns out it’s more complicated than that.
A parallel layer of communication
Neuropeptide signaling operates on slower timescales and can affect neurons beyond the reach of synaptic wiring. If we think of cell adhesion molecules as determining who can wire to whom, neuropeptide ligand-receptor pairs determine who can signal to whom through the extracellular medium. The combinatorics are rich: with dozens of neuropeptide families and their receptors, neurons can express different “chords”—combinations of ligands and receptors that define their participation in overlapping signaling networks.
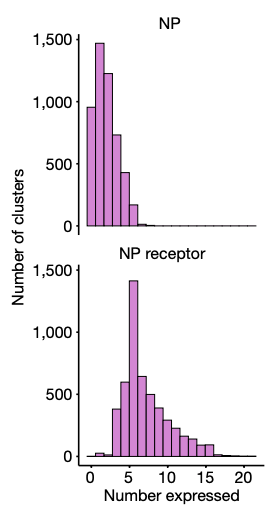
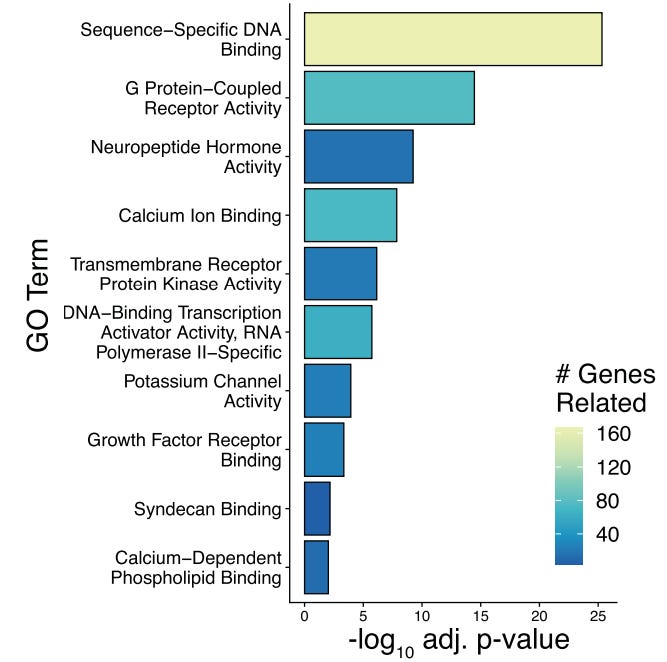
Strikingly, both the Allen and Broad Institute transcriptomic atlases found that neuropeptides are a major contributor to splitting transcriptomic clusters (Yao et al., 2023; Langlieb et al., 2023). What distinguishes one hypothalamic cell type from another at the molecular level is often which neuropeptides they make and which neuropeptide receptors they express. This suggests that functionally distinct neuron types are differentiated by neuropeptide “chords”, particularly in the Steering Subsystem where bespoke circuits require precise signaling specificity.
Integration-by-precise-wiring vs. integration-by-soup
You can see these very different methods of building circuits dramatically illustrated by line attractors. In dynamical systems, line attractors are characterized by a tight line of attractors—a line of marginal stability. Position along this attractor line tracks a single variable, and the attractor naturally remembers this variable over time, and so are ideal to accumulate evidence. They’ve been implicated in evidence accumulation in decision-making tasks, to track a slowly decaying variable like aggression or sexual receptivity, and in machine learning, to track complicated judgements about whether a sentence has positive or negative sentiment.
You have to work hard to make a good line attractor with wired connections. Khona & Fiete (2022) present one template:
Two neuron groups with in-group excitation and across-group inhibition, precisely tuned interaction strengths and quasi-linear neural input–output responses can counteract activity decay in the network and produce persistent activity over a continuum of activity levels in the two populations, defining ramp-like neural tuning and a line of attractor states. (emphasis mine)
Of course, it’s not impossible to learn a line attractor—RNNs trained to accumulate evidence will learn line attractors—but it requires precise tuning.
By contrast, building a line attractor is trivial to build with neuropeptides. Neuropeptidergic signaling provides a natural mechanism for analog integration—the slow accumulation of signals over time. When neurons release neuropeptides into the extracellular space, those molecules persist far longer than the millisecond timescales of synaptic transmission. A population of neurons releasing the same neuropeptide create a shared pool that integrates their collective activity; neurons expressing the receptor effectively “read out” this integrated signal. I call this process of analog integration of diffusable signals in the extracellular space integration-by-soup (I hope it catches on).
Integration-by-soup is a less flexible mechanism than integration-by-precise-wiring: diffusion constants are hard-wired; it can cause interference across space. For things that relate to survival, however, they can be just the right mechanism, provided that there is a fixed number (~100) of slowly changing state variables that need to be tracked.
The VMH Line Attractor: A Case Study
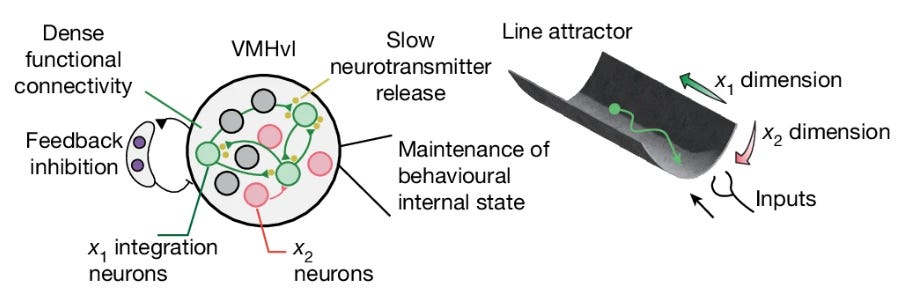
A hypothalamic circuit that tracks aggressivity illustrates how wired and wireless mechanisms could work together to create a line attractor (beautifully reviewed here). Nair et al. (2023) first described neurons in the ventromedial hypothalamus (VMH) expressing estrogen receptor 1 (Esr1). These neurons encode an aggressive internal state during both fighting and observation of fighting: dynamical systems analysis revealed an almost perfect line attractor in state space, with very slow temporal decay (>50s). In Vinograd et al., 2024, they identified two functionally distinct subpopulations within this genetically defined cell type:
x₁ neurons form the “integration dimension”—they show slow dynamics with long time constants, accumulating activity during aggressive encounters. Activity projected onto this dimension persists after stimuli are removed, ramping up along a line attractor that correlates with escalating aggression from sniffing to mounting to attack.
x₂ neurons form an orthogonal dimension with faster dynamics. These neurons show transient responses time-locked to stimulus onset but do not integrate over time.
The critical finding concerns functional connectivity: single-cell optogenetic stimulation combined with calcium imaging revealed that x₁ neurons are functionally interconnected with each other but not with x₂ neurons. When x₁ neurons are stimulated, other x₁ neurons show increased activity (functional connectivity within the ensemble), but x₂ neurons do not respond. Conversely, stimulating x₂ neurons produces only transient off-manifold perturbations, and the neural trajectory returns to the line attractor rather than integrating.
Important technical distinction: functional connectivity between x₁ neurons means that when some x₁ neurons are zapped, others activate; it does not say anything about whether they’re directly connected, indirectly connected, or wirelessly connected. You would need electrophysiology and/or connectomics to infer anatomical connectivity.
Two ways to connect: choose one
The circuit could either be mediated by some bespoke wired circuitry or some bespoke wireless circuitry. Let’s look at them one by one. In the Barabási model, this circuit might arise from biclique operators:
All VMHvl Esr1+ neurons share a common transcriptomic identity that positions them in the same nucleus and allows them to form synapses with each other.
Within this population, additional transcription factors or cell surface molecules distinguish x₁ from x₂ subpopulations. If x₁ neurons express a specific adhesion molecule or receptor that x₂ neurons lack, a biclique operator can generate exactly the dense within-x₁ connectivity observed.
But there’s another way of doing this, this time wirelessly: if all the x₁ cells express some neuropeptide receptor, they could accumulate aggression information from upstream sensory neurons via integration-by-soup. So which one is it?
It seems in this case that it’s wireless integration-by-soup. In companion work, Mountoufaris and colleagues showed that knocking out neuropeptide signaling (OXT and AVP receptors) in VMHvl neurons eliminates the line attractor dynamics (Mountoufaris et al., 2024). However, the authors note that “that does not exclude a contribution from recurrent glutamatergic excitation in the ventromedial hypothalamus, as in line attractors that mediate cognitive functions on shorter time scales”. In other words, it could actually be a little bit of both, but it definitely looks like neuropeptides are involved.
I don’t want you to come off with the impression that neuropeptides are the source of all line attractors in the brain; far from it! Line attractors are ubiquitous, and those that are involved in e.g. decision-making and evidence accumulation in the cortex are likely done through conventional wiring. However, especially in parts of the brain that are part of the steering subsystem, neuropeptides can do computational heavy lifting beyond their reputation as neuromodulators.
This example illustrates the deep connection between cell types, connectivity, and computation. The genome doesn’t specify “build a line attractor for aggression.” It specifies molecular identities (Esr1+, plus whatever distinguishes x₁ from x₂) that, through wiring rules and neuropeptide channels, generate the connectivity topology that implements line attractor dynamics.
More epicycles
I presented a binary story of wired vs. wireless connectomes. Of course, the real story is far more complex, with the gnarliest complication being space. Distance strongly constrains which neurons connect with which. Layout is ultimately encoded in the genome; and the genome dictates axon guidance, through different guidance molecules like netrins, slits, semaphorins, and ephrins. That means transcriptomic identity also influences layout and guidance signals. it could thus be said that cell types influence connectivity through (at least!) three interacting means: local wiring, wireless neuropeptides, and global layout.
Not only that, but these factors are dynamic! Neurons migrate during development, which means that local connections can translate into long-range connections. Transcriptomic signatures also change during development. It’s possible, for example, that CAMs are expressed only at an intermediate timepoint during development, to determine connections, then go away during adulthood. You can see hints of this in the Yoo et al. (2023) data, which captured transcriptomes at different time points.
…I could continue all day. The take-home here is that there are myriad routes through which cell types influence connectivity, that the connection rules don’t appear particularly mysterious, and that studying cell types and connectomes at the same time is fruitful.
Conclusion
The question “how does a genome specify brain circuits?” has an elegant answer: through cell types. Combinatorial gene expression patterns determine surface molecule profiles; these implement wiring rules specifying who connects to whom; and connectivity topology determines computational dynamics. The genome encodes not just anatomy but function. They also determine neuropeptide patterns, creating a parallel channel of communication that we’re just starting to investigate.
This helps explain why, for example, the hypothalamus harbors over a thousand cell types, while the cortex tiles the same canonical circuit across vast territory. Innate behaviors require specific circuits, and specific circuits require specific cell types, which determine wiring and wireless rules. Studying these signatures could help us finally understand how evolution wrote the instructions for building a brain. Moreover, they could help us understand the rich tapestry of innate drives and behaviors that define different species.