Cell types: encoding the brain's BIOS
Inferring the structure of primary rewards from connectomics
In an interview late last year on the Dwarkesh Podcast, Adam Marblestone—CEO of Convergent Research, former research scientist at Deepmind, and a true polymath in connecting neuroscience and AI—discussed what insights we might extract from complete brain wiring diagrams. His central claim is that the most valuable information in a connectome concerns innate reward functions and motivational systems: the circuits that tell learning systems what to learn. Adam mentioned that one way we would know that an area of the brain specifies reward information is that it contains many different cell types.
Dwarkesh seemed very confused by this, asking a few different times: “Why would each reward function need a different cell type?” I empathize with Dwarkesh here! It is mysterious that a cell type could represent something as abstract as a reward. As a computational neuroscientist who mostly worked at the representation level during my PhD, I’ve leaned historically towards thinking of cell types as a mere “implementation detail”. But over conversations with Adam, Steve Byrnes, Paul Cisek, Tony Zador, and a few others, I’ve started to become convinced that cell types are a really useful lens to think about innate behaviors and rewards.
In this essay, I’ll unpack the conversation and answer the question: what do cell types have to do with reward functions? To answer it, we’ll need to understand what kind of information can be encoded in the genome, and how that information ultimately relates to connectomes and to cell types. I’ll connect the answer to the central claim of Adam: that these connections matter for AI, and AI safety in particular.
Some things are innate
Andrew Barto and colleagues make the point that all primary rewards are internal, and must be genetically encoded. In reinforcement learning, which Barto co-developed along with Rich Sutton, an agent learns by receiving reward signals that indicate what is good and bad. The critical insight is that for biological organisms, all of these reward signals are internal—they are generated by the organism’s own nervous system. It is not a chunk of steak that gives reward: it is circuitry inside the brain that assigns positive valence to fat, salt, umami, heat, and texture. Things like money—secondary rewards—must be bootstrapped off of the pre-existing primary rewards.
The foundational, primary reward signals must be written into the genome. Now, you might ask, can’t we learn what is rewarding by imitating our parents? Ah, but why would we find it rewarding to imitate our parents in the first place? At some point, the ladder of rewards bottoms out to a foundational structure: a kind of BIOS of the brain. The genome can’t literally specify exact connections of the brain, however; the roughly 3 billion base pairs are dwarfed by the approximately 100 trillion synapses in the brain. This mismatch, what Tony Zador calls the “genomic bottleneck”, means that evolution must compress circuit specifications into a limited information channel. Rather than encoding individual wires, the genome encodes wiring rules—and cell types are what make these rules executable.
Innate behaviors are encoded by cell types
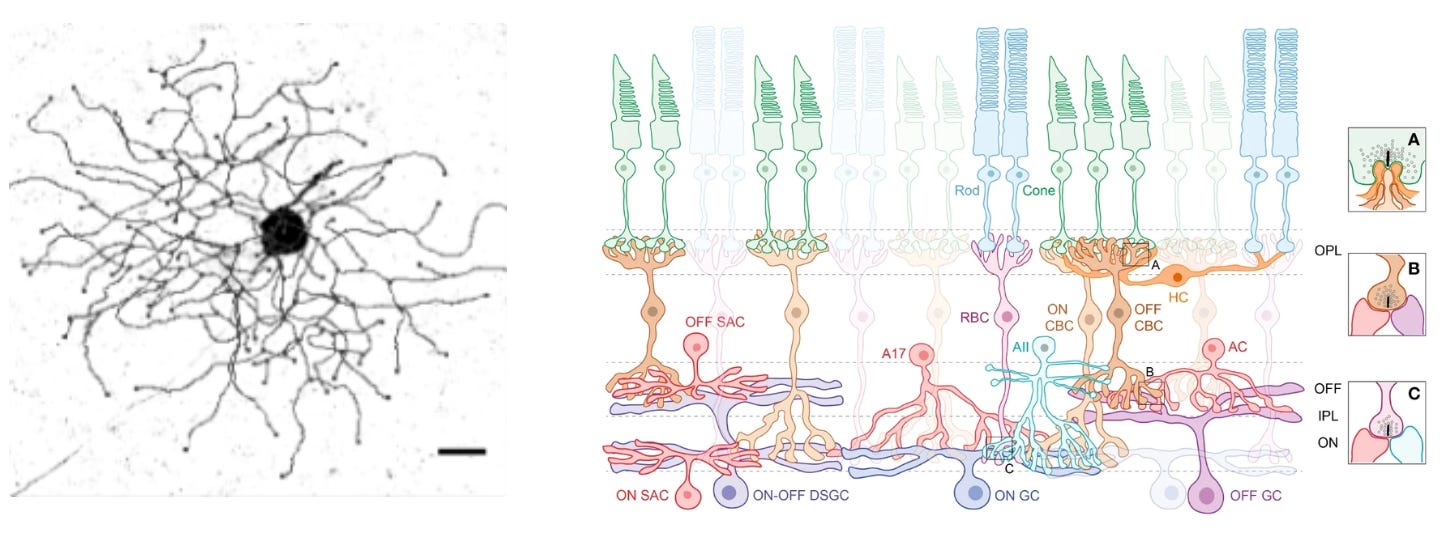
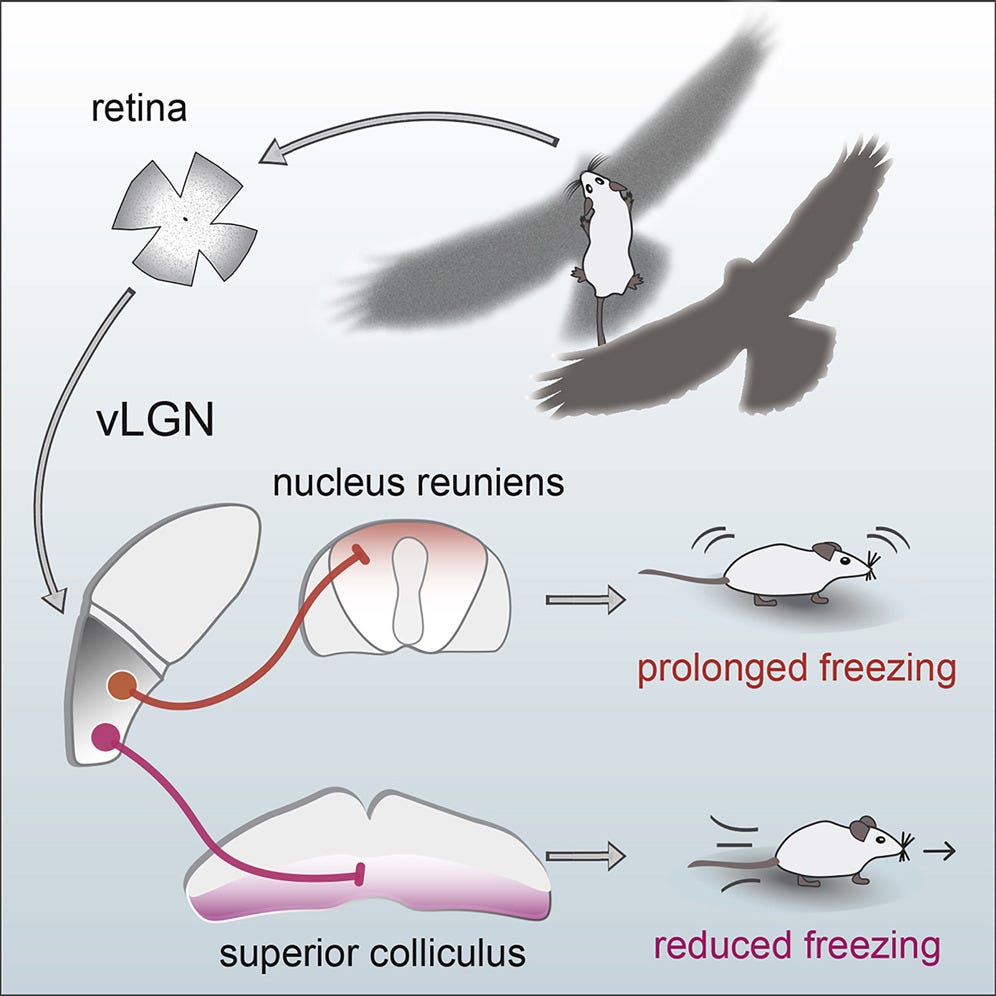
Let’s build some intuition about how cell types can specify an innate behavior. Mice live in fear of hawks and owls. Having the right instincts when they encounter flying predators can mean the difference between life and death. In their retinas, mice have a very peculiar and rare type of retinal ganglion cell—OFF-transient alpha RGC (tOFFα)—that is selective for dark looming things in the upper half of the visual field (e.g. scary birds). These cells are causally implicated in a reflex: mice either freeze in a prolonged manner or escape when they see a dark looming stimulus overhead.
The retinal ganglion cell’s role is determined both by morphology (shape) and connectivity. This cell has a large dendritic arbor that stratifies in the right layer of the retina to receive input from specific amacrine cell types. It’s connected downstream to the right defensive behavior circuits— among others, the superior colliculus. Hence, both morphology and connectivity relate to its functional role.
Here’s a key idea: if you look at an area of the brain, and it has many idiosyncratically shaped and connected neurons, that probably means that area encodes innate behaviors and rewards. Whereas if you go in an area and you find a quasi-crystalline, repeating structure with lots of similar neurons (e.g. in the cerebellum or cortex), that probably means that it derives its role by learning from the environment.
There are many ways of defining cell type, the canonical one is now transcriptomic
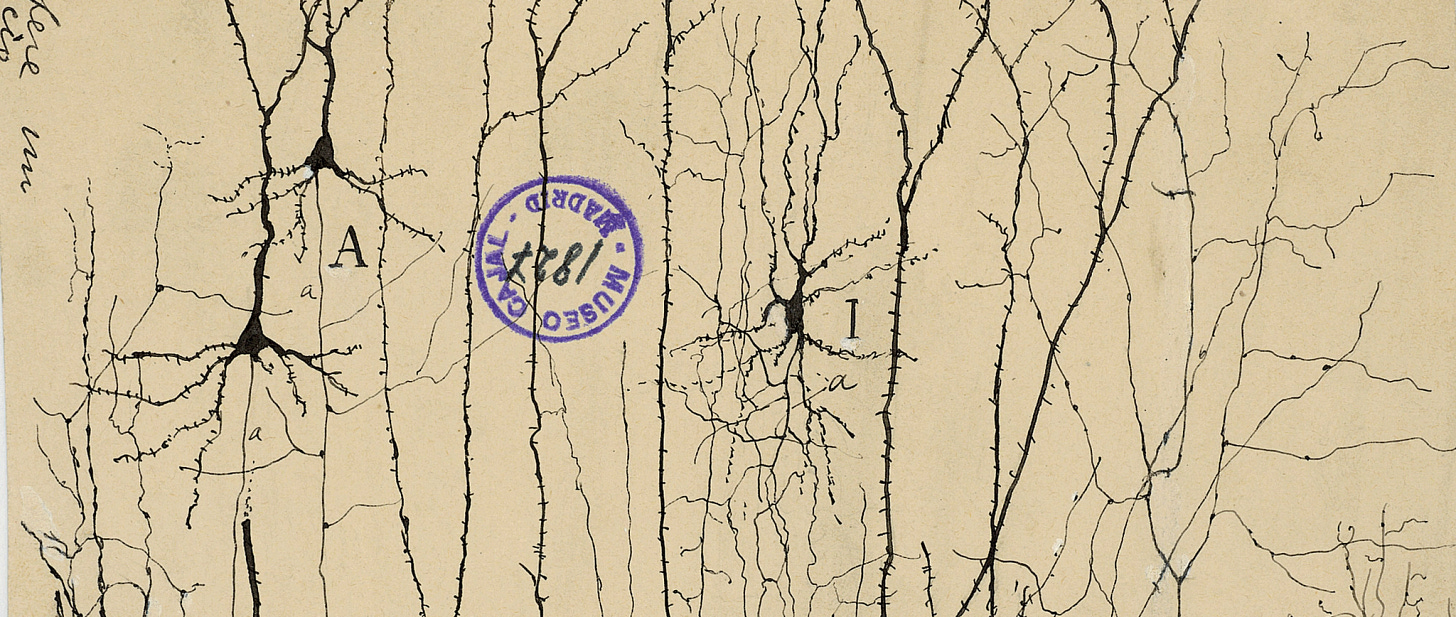
I just outlined one way that you could find putative learning vs. innate areas in the brain: count the weird cell types. But cell types are one of those seemingly precise biology terms that are hopelessly overloaded. Classically, cell types were determined exclusively by morphology—think Cajal’s beautiful tracings of nervous cells. Now that connectomics has started to mature, cell types are also increasingly defined by connectivity: two cells that share similar connectivity roles in a circuit are presumed to belong to the same cell type (see my previous piece on FlyWire).
But perhaps the most all-encompassing definition of cell type relates to cell fate. During embryonic development, neural progenitor cells undergo a series of fate decisions, progressively restricting their potential until they differentiate into a specific neuronal (or glial) identity. These fate decisions are locked in by epigenetic modifications and the stable expression of master transcription factors. Those expressions ultimately determine both shape and connectivity (more on this later).
Transcriptomic identity is thus now commonly used for cell-type classification. Each neuron expresses a particular subset of the genome’s ~20,000 protein-coding genes. We can measure cell types by hierarchically splitting clusters based on their expression profiles. The genes that split the clusters include genes that relate to neurotransmitters, neuropeptides ligands and receptors, cell-adhesion molecules, as well as transcription factors. That looming-selective retinal cell I mentioned earlier can be differentiated from other retinal ganglion cells, for example, by how much it expresses the gene Kcnip2 (Wang et al. 2021).
Lots of transcriptomic cell types = areas that define innate behavior
Both the Allen Institute’s and the Broad’s spatial transcriptomic cell type atlases from 2023 identified thousands of cell types (Yao et al. 2023; Langlieb et al. 2023). These cell types are not evenly distributed in the brain. Certain areas, including the hypothalamus, midbrain areas—including the superior colliculus—and the medulla, express far more cell types per total number of cells than cortex or cerebellum.
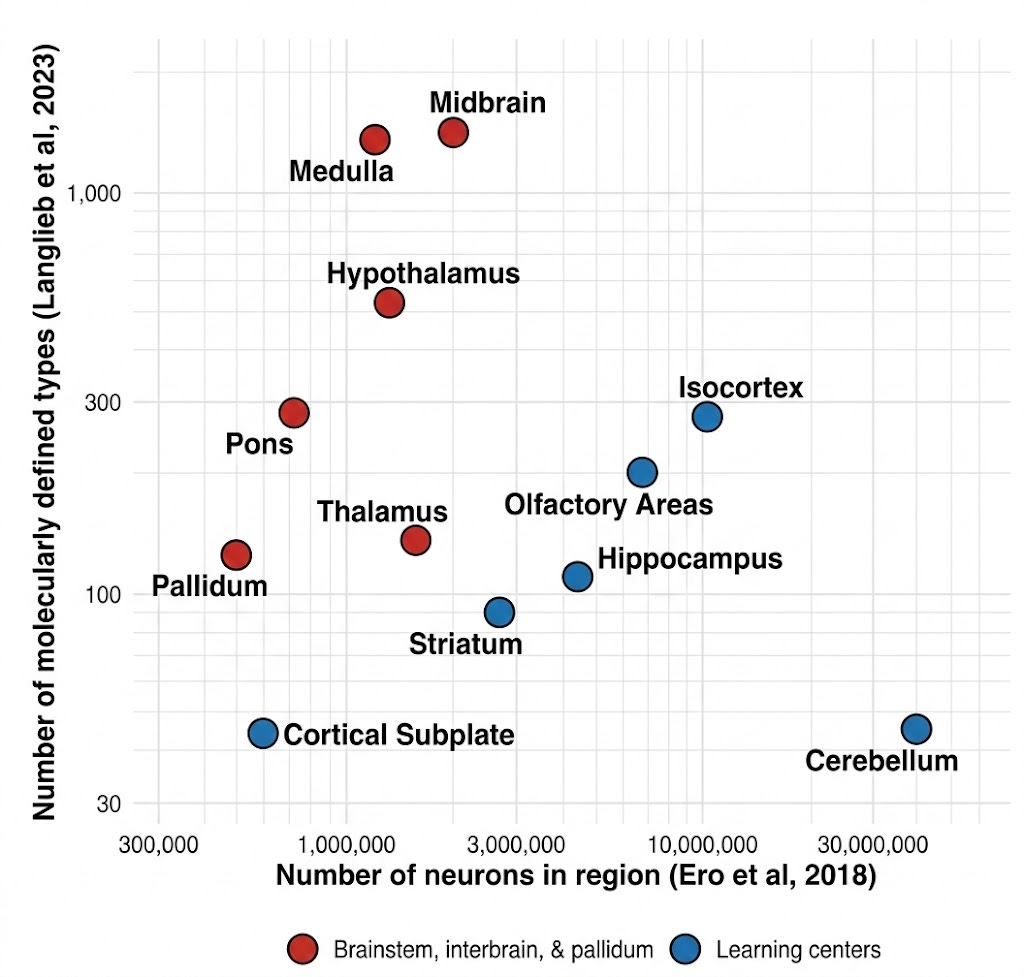
This makes sense. The hypothalamus regulates sleep, circadian rhythms, aggression, libido, hunger, etc. It keeps track of very important things for survival, which ought to be encoded in the genome. The superior colliculus (aka optic tectum) of the midbrain is the original visual structure. It orchestrates visual and motor responses. For instance, looming-selective retinal ganglion cells project to the superior colliculus to drive the freeze-or-flight response to overhead hawks.
Steven Byrnes predicted this distinction; the framework is now being adopted
This distinction between innate and learned systems had, in years prior, been framed by Steven Byrnes (Byrnes, 2022):
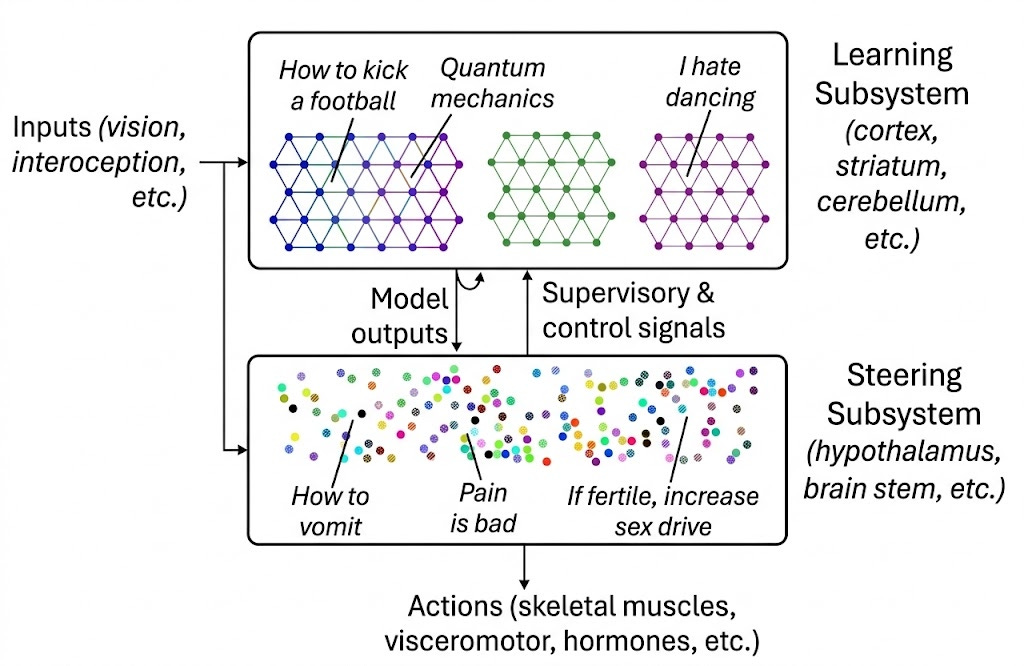
The Learning Subsystem—primarily the neocortex, but also the cerebellum—implement generic, general-purpose learning algorithms. It builds world models, learns sensorimotor mappings, and acquires skills through experience. The same basic circuit motifs—in cortex, roughly six layers of neurons with stereotyped connectivity—are tiled across areas devoted to vision, language, motor control, and abstract reasoning. What differs between areas is primarily what inputs they receive and what outputs they produce, not their fundamental computational architecture.
The Steering Subsystem—comprising the hypothalamus, brainstem, amygdala, and related structures—is entirely different. It implements bespoke, artisanal circuits: hand-crafted by evolution over millions of years, each one tuned to detect specific stimuli or generate specific behaviors. When you feel hungry, afraid, lonely, or curious, these signals originate in the Steering Subsystem. Each distinct motivation—salt appetite, fear of heights, sexual attraction, maternal care—requires dedicated circuitry that cannot be learned from scratch.
It’s pretty remarkable that Steven—an AI safety researcher, not a neuroscientist by training—accurately predicted this split prior to the publication of large-scale transcriptomic atlases. Adam, during the podcast, goes on at length about some of the more subtle aspects of his framework. It is just starting to gain traction along neuroscientists, including in a recent, lucid editorial by Fei Chen and Evan Macoscko, the PIs from the Broad Institute who published one of the original whole-mouse-brain transcriptomes.
In their editorial, they frame the learning and steering subsystems as two radically different ways of wiring up a brain, with very different scaling laws: one made by piling up heuristics passed down through generations; and the other made from a very good, generic learning algorithm. It will not escape some of you that the learning subsystem has been vastly elaborated in mammals, and in primates in particular. Indeed, one could argue that this was evolution’s “bitter lesson” moment: switching from a pile of carefully tuned heuristics to a generic learning system. Yet, it is the interplay between these two systems that makes learning possible; for example, the superior colliculus never disappeared despite the visual cortex being vastly elaborated in primates.
If you’re going to do connectomics, the steering system is a good place to do it
Adam makes the point in the podcast that connectomics may be most useful for understanding the steering system. The steering subsystem is stereotyped; its structure tells you something about intrinsic behaviors and circuits. By contrast, the connectome of a cortical column—as was done in the MICrONS project, for example—represents a static snapshot of a dynamic process. The unique thing that is ascribed to cortex—learning—is quite possibly invisible if you’re just looking at static connections. Looking at cortex at the learned representational level makes more sense.
In my mind, the connectome of the fruit fly is quite useful precisely because fly brains are highly stereotyped—though they show some learning, e.g. in the mushroom body. The bespoke circuits in the steering subsystem of mammals are of the same nature. As Adam mentions, they are probably hiding insights not just about instinctual behaviors and homeostasis, but also primary rewards. Human rewards are built from the distillation of millions of years of evolution; it’s what allows us to maintain adaptive behavior over a broad range of circumstances.
Those rewards are akin to a finely tuned, ten-thousand-line Python file full of if-elses that specify behavior, not an elegant deep net that Karpathy would dream up. We know that misspecifying rewards in real RL systems leads to various types of reward hacking, from missing the point of a game, to staring at noisy TVs, to sycophancy and slop. Humans are mostly robust to this. Inferring our rewards from behavior is the subject of inverse reinforcement learning (IRL), and it is very ill-posed problem. If we could instead decompile the circuits in the steering subsystem to that ten-thousand-line Python file, we would be in good shape to create aligned AI systems. At the heart of this decompilation effort is a map of the steering subsystem—a connectome. This is easier said than done, but it is clearly one of those “big if true” ideas: a reward function, written in a language that we could inspect and potentially port to artificial systems.
This suggests that the right target for AI alignment may be these deeply ingrained reward circuits, our motivational bedrock. Pain avoidance, social bonding, curiosity, care for offspring—these aren’t learned preferences but genome-specified circuits that learning systems are built to serve. The Steering Subsystem’s bespoke circuits could serve as a template for building AI that shares our sense of what matters; but first, we have to map it.
In the second part of this series, I explain how, mechanically, neurons of different cell types recognize each other to build bespoke circuits. Read it here.



Restacked, excellent piece! I've had in mind the metaphor of "firmware" for the lower brain and now you have me wondering which of that and BIOS minimizes muddle (all metaphors are wrong! but some are useful). Besides NeuroAI, I believe medicine itself (including specialties beyond psych/neuro/neurosurgery) may benefit greatly from decompiling the lower brain. Let's keep pushing on this from all angles.
Excellent and insightful read as always. Curious, to your mind, to what extent can we think of an AI's default system prompts/system instructions as a sort of analogue to this basal "steering system" in biological brains? Seems like we already do conceive of it this way in LLMs, with the weights of the model itself being the thought forms of the cortex (generalizable, emergent, acquired through extensive RL), and the default instructions being what the model is "born" with? I suppose the analogy breaks down when you consider that the system prompts instruct a lot of higher-level concepts like model identity or tonality.